
David Nasonov
MSc Applied Bioinformatics Graduate looking for work
About Me
Hi! Welcome to my website, I am an AI-researcher with expertise in biologically inspired machine learning and bioinformatics.
Currently building Insular AI, where I've already engineered ML models for autonomous systems achieving 25% higher performance than reinforcement learning with 10× lower inference cost.
MSc Applied Bioinformatics (King’s College London), specialising in mass spectrometry pattern recognition (Python, TensorFlow, Pandas). Contributor to the Seurat GitHub repository (2.5k★), where I improved usability for single-cell RNA-seq analysis.
First-class BSc Biotechnology graduate and Lonza Biotechnology Prize winner.
When I'm not bashing my head trying to debug a piece of code, I can be found competing (and often losing) in grappling
Experience
Co-Founder / AI researcher
Insular AI (Stealth) - London, UK
• Engineered bio-inspired ML models achieving 45% higher task success rates vs reinforcement learning, with 5x lower training costs and 10x faster inference. • Developed multi-framework implementations (Flax, JAX, PyTorch), ensuring reproducibility and scalability across platforms while coordinating code integration through GitHub. • Secured early investor interest by producing concise pitch materials and data-driven visualisations that translated technical innovation into commercial value. • Led collaboration with a 5-person team, incl. a King’s College London professor, to design and benchmark novel bio-inspired architectures supporting interoception and continuous learning in autonomous systems.
Treasurer
Russian Speaking Society - Nottingham, UK
• Through campaigning for membership purchase and introducing member-funded events, I increased the budget by 36%. • Spearheaded the launch of flagship events, including interviews with renowned journalists like Oleg Kashin.
Research Placement
University of Nottingham - Nottingham
• Analysed 96 single-cell RNA-seq datasets from three pluripotent stem cell types, identifying 23 candidate genes regulating pluripotency. • Built a Boolean Gene Regulatory Network with PyBoolNet, modelling three different stages; adopted in an MRC grant proposal. • Delivered publication-quality visualisations (volcano plots, PCA, heatmaps) to support findings and a stronger grant application • Applied differential gene expression, GO-term and pathway analyses to map regulatory mechanisms, integrating results with an extensive literature review. Under Prof. Dov Stekel
Go to Market Consultant
SnazzyBird - Nottingham
• Lead a multidisciplinary group of 5 students working on a consultancy project for a client in sustainable furniture. • Collaborated with the team to schedule and organise weekly meetings between multiple stakeholders. • Implemented MoSCoW prioritisation technique using a Trello board for managing tasks effectively. • Completed the project ahead of schedule, increasing the B2B reach by 40%.
Featured Projects
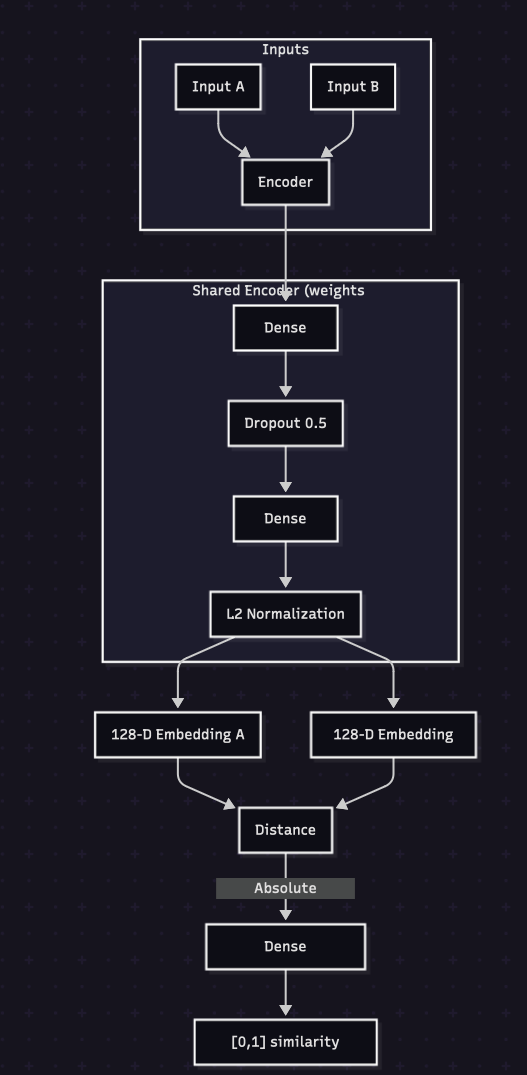
MSc Applied Bioinformatics Dissertation
Part 1: Developed an aggregate model, combining results from 3 unsupervised learning algorithms (Iso Forest, DBSCAN, Autoencoder) to flag athletes suspected in growth hormone abuse based on biomarkers Part 2: Built a Liquid Chromatography - Mass Spectrometry raw data pre-processing pipeline. Used the pre-processed data to train a Siamese Neural Network model to identify if two samples come from the same person or not. AUC = 92.3%, F1-score = 0.91
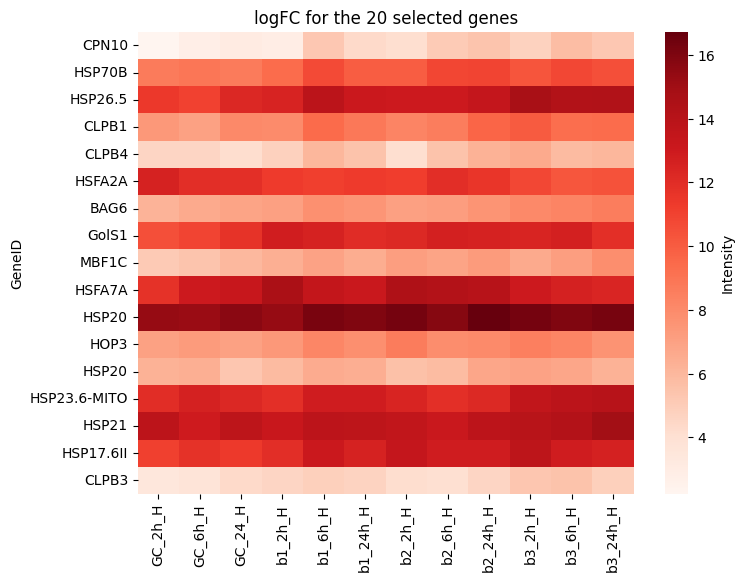
BSc Biotechnology Dissertation
Heat stress during flowering poses a significant threat to oilseed rape (Brassica napus) production, necessitating a comprehensive understanding of the molecular mechanisms underlying plant responses to heat stress. Oilseed rape is a globally important crop with diverse applications, including as a source of vegetable oil, animal feed, and biofuel. However, its susceptibility to heat stress presents a challenge to sustainable cultivation. In this study, the genetic factors mediating heat stress response during flowering in oilseed rape were investigated. Here we show that heat shock factors (HSFs) and heat shock proteins (HSPs) play pivotal roles in the plant's ability to withstand heat stress. Complementing previous studies in other species, our findings reveal the specific upregulation of HSP20-like chaperones superfamily proteins, CLPBs, GolS1, BAG6, HOP3 and MBF1C inn response to heat stress, underscoring their potential as targets for genetic modification to enhance heat tolerance in oilseed rape. These results lay the groundwork for further study of heat stress response in Brassica and the potential development of genetically engineered varieties with improved heat tolerance. By elucidating the genetic and molecular basis of heat stress response in oilseed rape, this study has broader implications for sustainable crop production in the face of escalating environmental challenges associated with climate change.
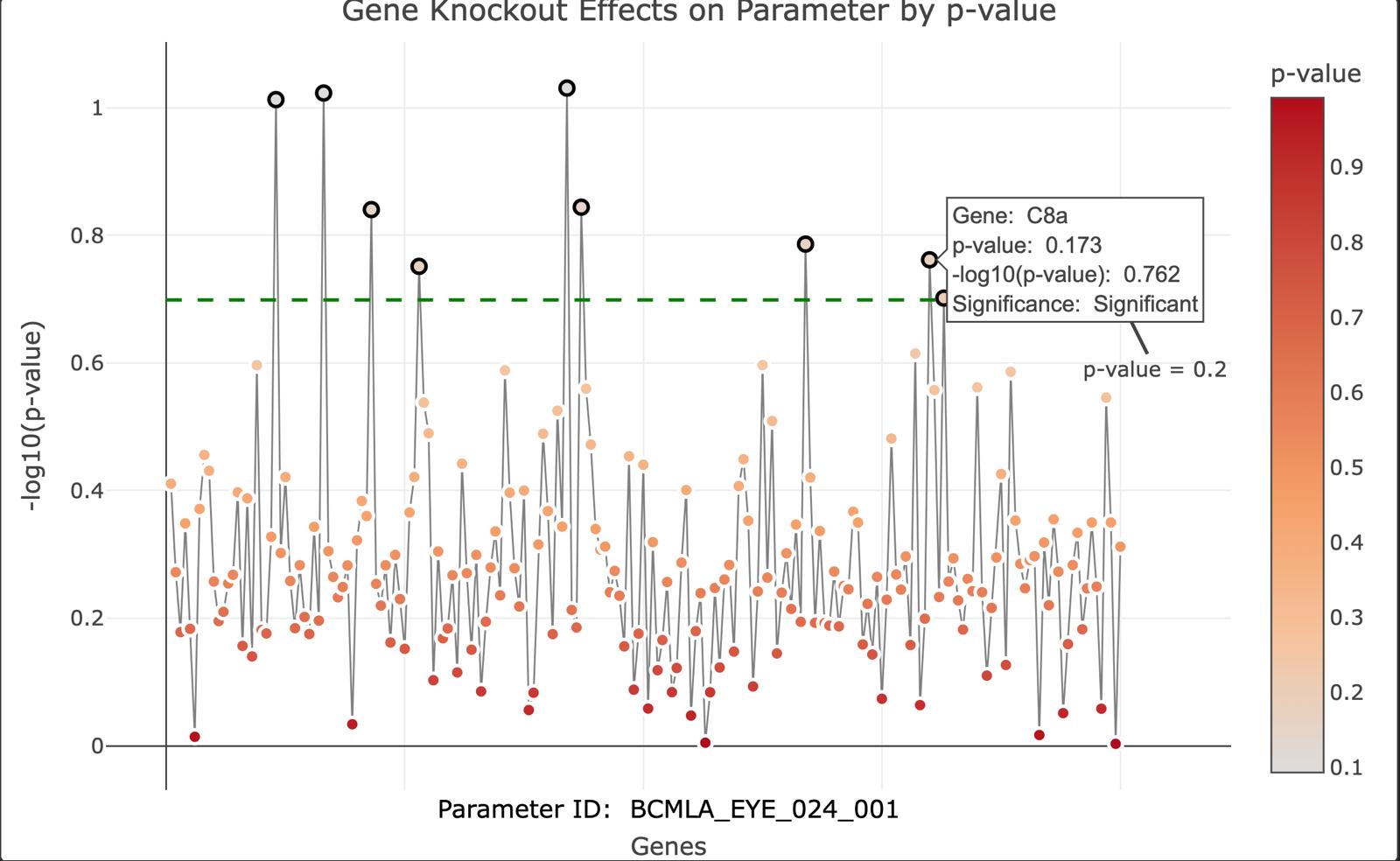
Mouse Phenotyping data analysis
- Raw data from the IMPC, in the form of 180,000 individual csv files, containing information about a single analysis, assessing effect of knocking out a gene on a given phenotype parameter was collated and cleaned - Parameters were grouped to reduce parameter space (275 -->19) - A scalable and comprehensive SQL database was created to efficiently store and query the data(Architecture seen on GitHub) - R Shiny app created to visualise a) the effect of the knockout every genes on a parameter, and the effects of knocking out one particular gene on every parameter ( Figure below and more on the GitHub page) - Project received 85% mark
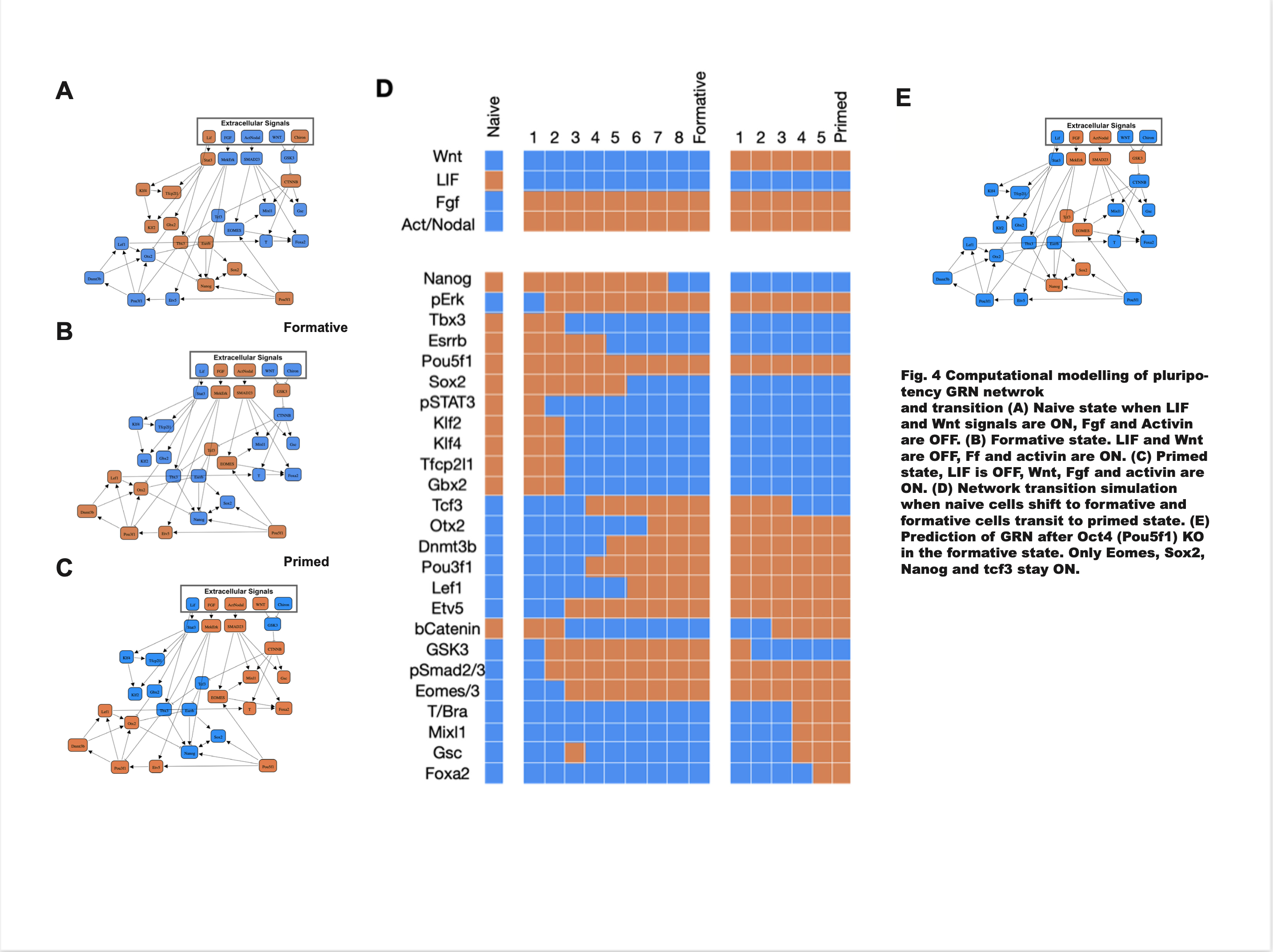
Boolean Gene Regulation Network
- Processed the data obtained from single cell RNA-Seq of three different types of pluripotent stem cells in mice (raw counts) - Visualised the data, using volcano plots, PCA and heatmaps (seaborn, matplotlib) - Downstream analysis including Differential Gene Expression, GO-term analysis, pathway mapping (kegg/reactome) was performed in python and bash - Using the results from the analysis, and the existing literature, key set of genes for pluripotent differentiation and their relationships were determined - A Boolean GRN was constructed, where by manipulating the presence or absence of 5 extracellular signals, the gene activation state was modelled accurately
Let's Connect
I'm always interested in discussing new opportunities and exciting projects.
Email: davidnasonov@gmail.com